
Research
Graduate Student

Abstract
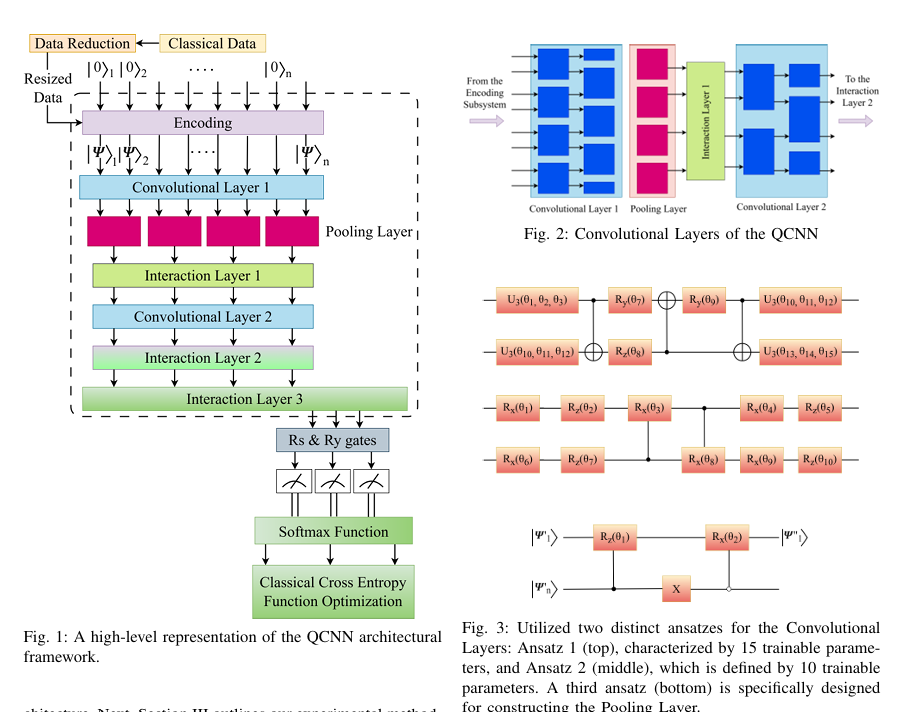
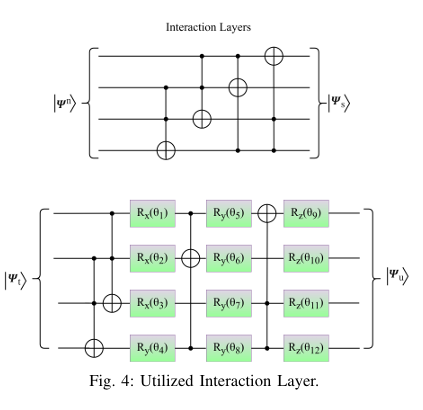
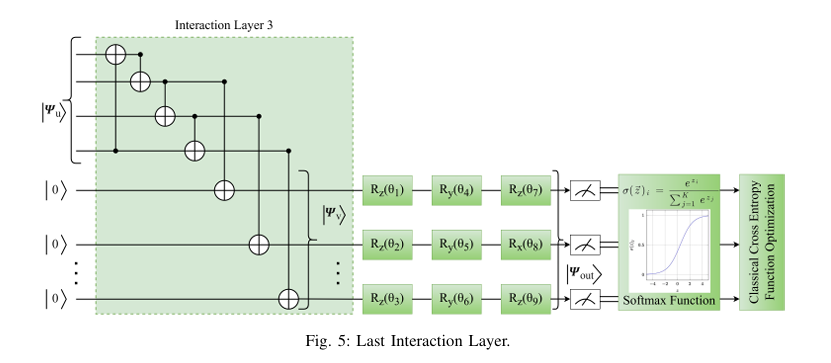
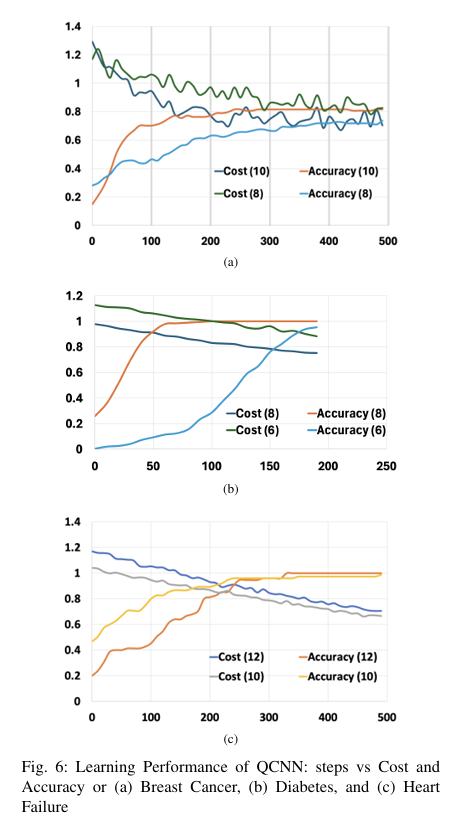
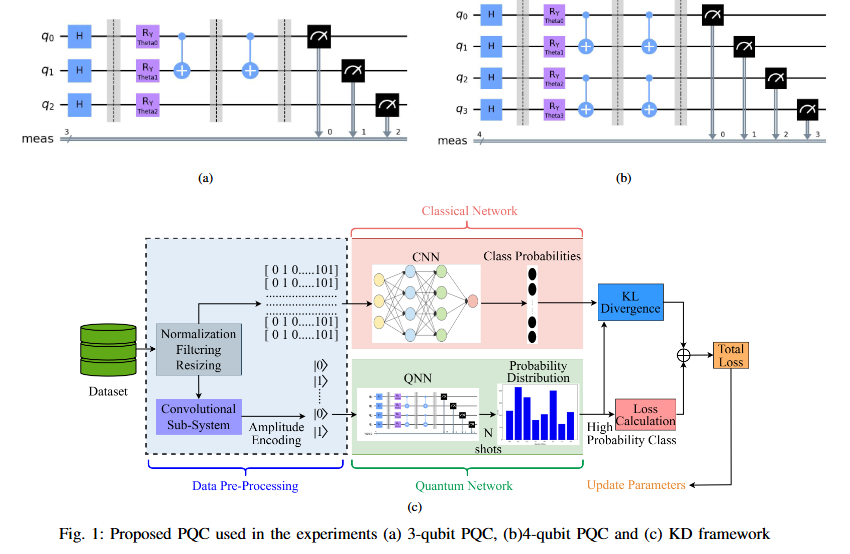
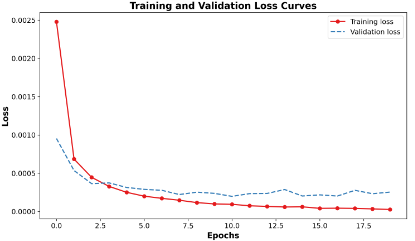
Quantum computing holds considerable promise for artificial intelligence (AI) in clinical decision support systems (CDSS), particularly in resource-constrained environments. This paper investigates a multiqubit quantum convolutional neural network (MQ-CNN) for medical diagnostics, leveraging param eterized quantum circuits to process low-resource healthcare datasets. We evaluate the framework on three binary classifica tion tasks: breast cancer (Wisconsin dataset), diabetes (Pima In dians), and heart failure prediction, using angle-encoded clinical features. The MQ-CNN achieves test accuracies of 82.4%, 98.7%, and 97.3% respectively, matching classical CNNs while reducing trainable parameters significantly. Comparative analysis shows the quantum model converges 22% faster than hybrid quantum classical counterparts under identical training conditions. Ro bustness evaluations confirm ≤3% accuracy degradation when subjected to 15% synthetic label noise. These results highlight the architecture’s suitability for resource-constrained environ ments, demonstrating that quantum-enhanced feature extraction can maintain diagnostic accuracy while significantly reducing computational overhead. This work provides empirical evidence for near-term quantum machine learning in practical healthcare applications.
BibTeX
Click to copy@INPROCEEDINGS{11172247,
author={Ovi, Tareque Bashar and Bashree, Nomaiya and Alam, Ayat Subah and Tanzim, Rawnak and Wahed, Md Abdul and Nyeem, Hussain},
booktitle={2025 International Conference on Quantum Photonics, Artificial Intelligence, and Networking (QPAIN)},
title={Multiqubit Quantum Convolutional Neural Networks for Efficient AI-Driven Healthcare Analytics},
year={2025},
volume={},
number={},
pages={1-6},
keywords={Degradation;Decision support systems;Accuracy;Computational modeling;Qubit;Noise;Neural networks;Computer architecture;Feature extraction;Convolutional neural networks;Deep learning;QML;QCNN;feature interaction;qubit;health AI},
doi={10.1109/QPAIN66474.2025.11172247}}
Abstract
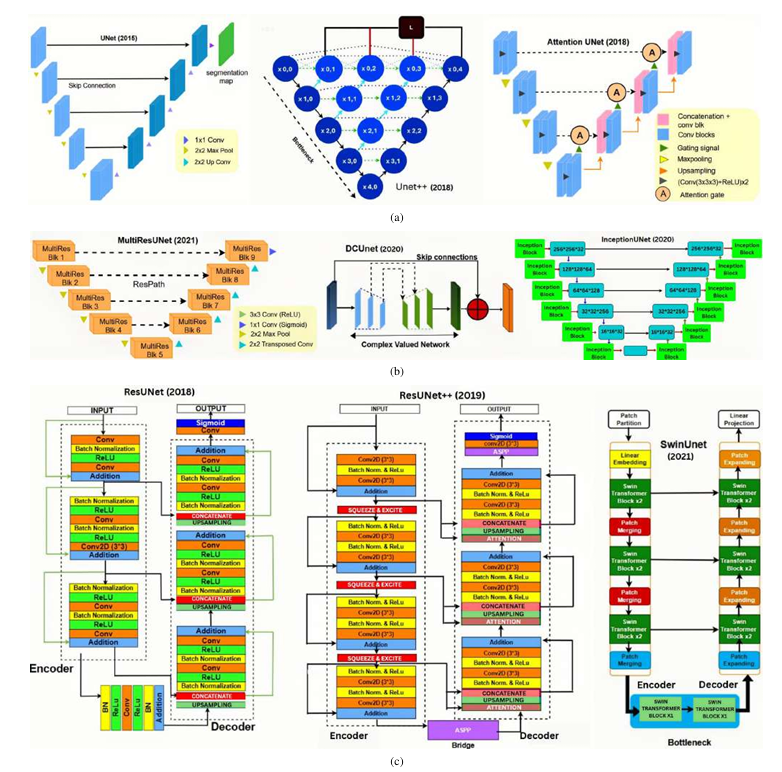
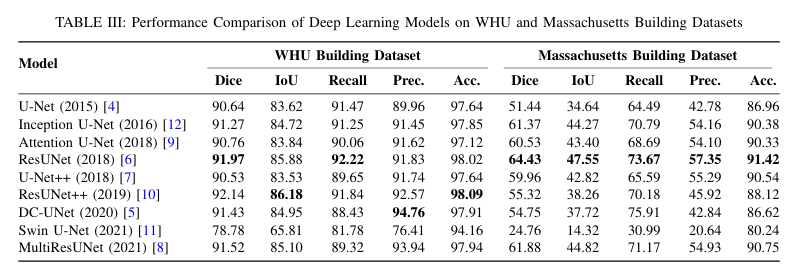
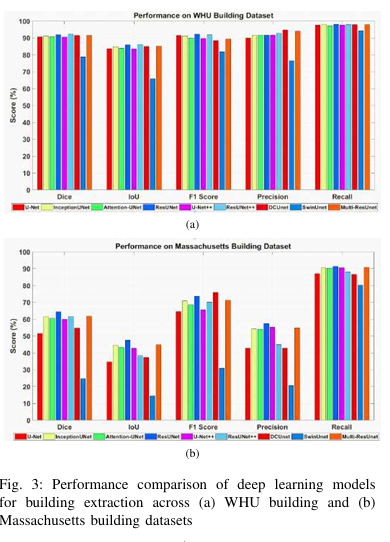
Cross-domain variations in satellite imagery-ranging from sensor characteristics to urban morphology-undermine the reliability of deep segmentation models. We isolate the architectural factor of residual connections by comparing nine encoder-decoder designs, trained from scratch on the large-scale WHU and the compact Massachusetts building datasets without pretrained backbones or patch augmentation. Residual architectures (ResUNet, ResUNet++) consistently lead by margin: up to 1.6 % higher Dice and 2.5% higher IoU on WHU dataset, and a 5% Dice gain on Massachusetts dataset. Error-mode analysis shows that skip-augmented identity mappings preserve feature gradients across depth, reducing false negatives by as much as 12 % versus vanilla U-Net under data scarcity. These results demonstrate that residual links are a simple yet powerful strategy to fortify building segmentation against domain shifts in real-world satellite imagery.
BibTeX
Click to copyS. B. Zahid, R. Tanzim, T. B. Ovi, N. Bashree, H. Nyeem and M. A. Wahed, "Impact of Residual Connections on Cross-Domain Generalization for Building Segmentation," 2025 International Conference on Quantum Photonics, Artificial Intelligence, and Networking (QPAIN), Rangpur, Bangladesh, 2025, pp. 1-6, doi: 10.1109/QPAIN66474.2025.11172159. keywords: {Training;Attention mechanisms;Architecture;Semantic segmentation;Buildings;Transfer learning;Transformers;Feature extraction;Satellite images;Remote sensing;building extraction;remote sensing;semantic segmentation;aerial imagery;attention mechanism;transformer;transfer learning},
Abstract
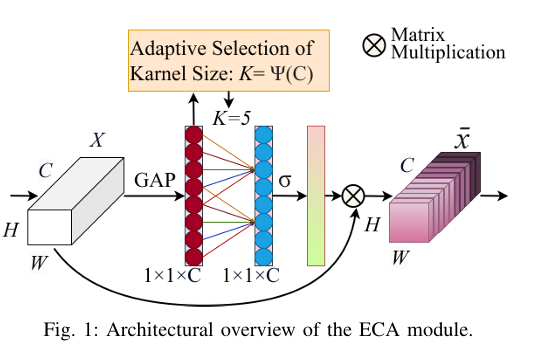
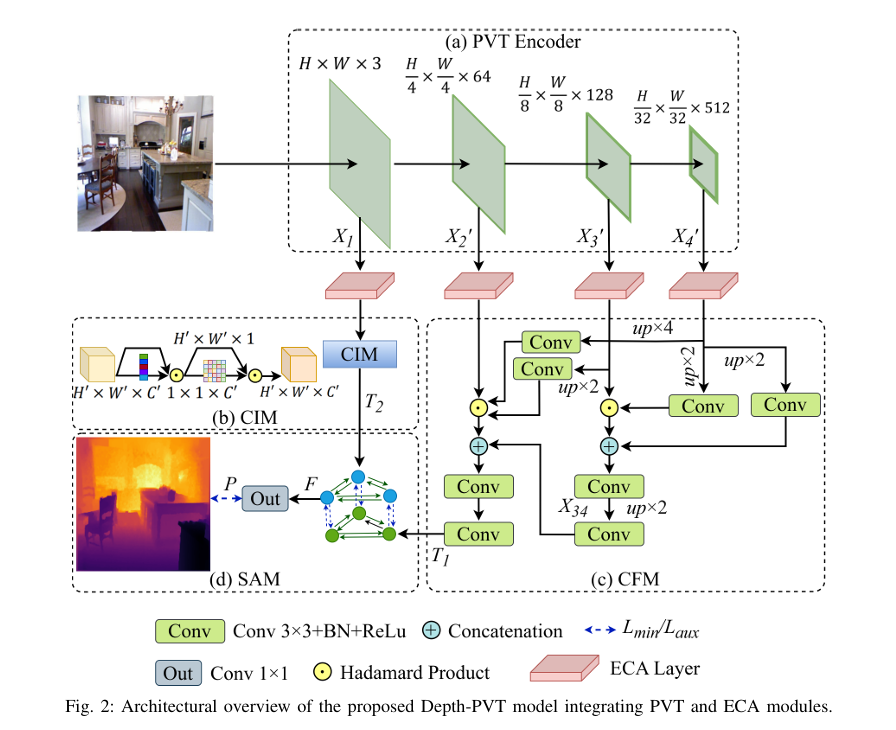
One of the most important machine vision tasks is monocular depth prediction, essential for applications such as autonomous navigation and augmented reality. While CNN-based architectures are widely used, they are limited by restricted receptive fields, which hinder effective multi-scale feature extraction. In order to overcome these issues, we develop Depth-PVT, an innovative architecture that integrates Efficient Channel Attention (ECA) with the Pyramid Vision Transformer (PVT). This integration enhances feature representation by emphasizing salient channel-wise features and suppressing irrelevant ones with minimal computational overhead. Depth-PVT incorporates three key modules within the PVT framework: the cascaded fusion module (CFM) for aggregating semantic and spatial information of high level features, the camouflage identification module (CIM) for capturing complex depth cues and low level features, and the similarity aggregation module (SAM) for fusing cross-scale features, thereby enriching depth d
BibTeX
Click to copy@inproceedings{ovi2025depth,
title={Depth-PVT: Pyramid Vision Transformer with Channel Attention for Depth Estimation},
author={Ovi, Tareque Bashar and Bashree, Nomaiya and Tanzim, Rawnak and Tirtha, Anindya Chanda and Nyeem, Hussain and Wahed, Md Abdul},
booktitle={2025 International Conference on Electrical, Computer and Communication Engineering (ECCE)},
pages={1--6},
year={2025},
organization={IEEE}
}
Abstract
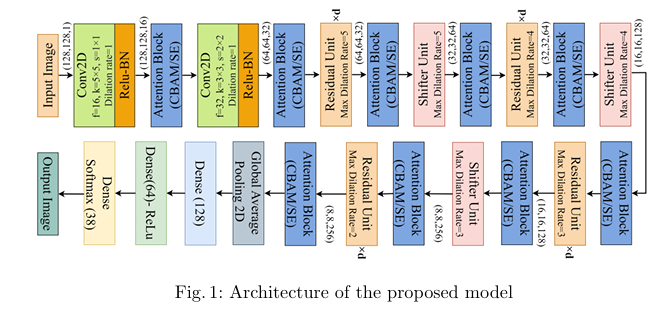
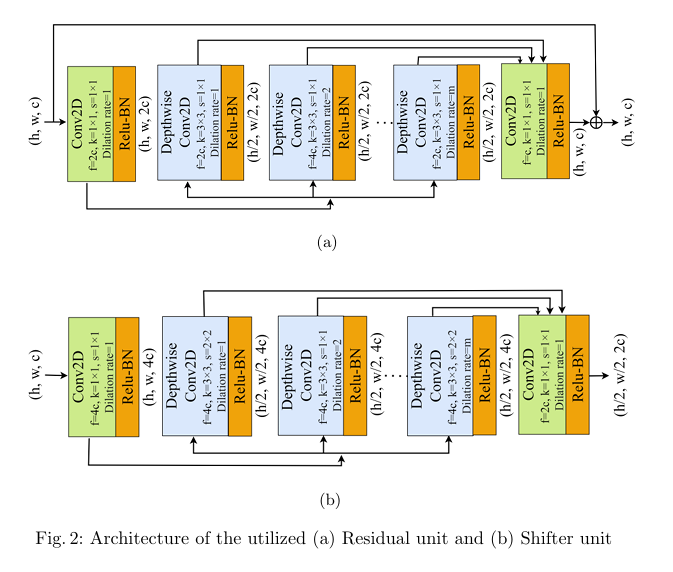
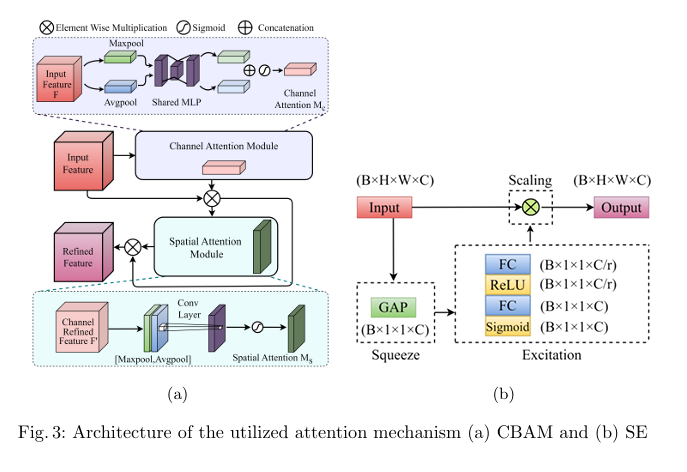
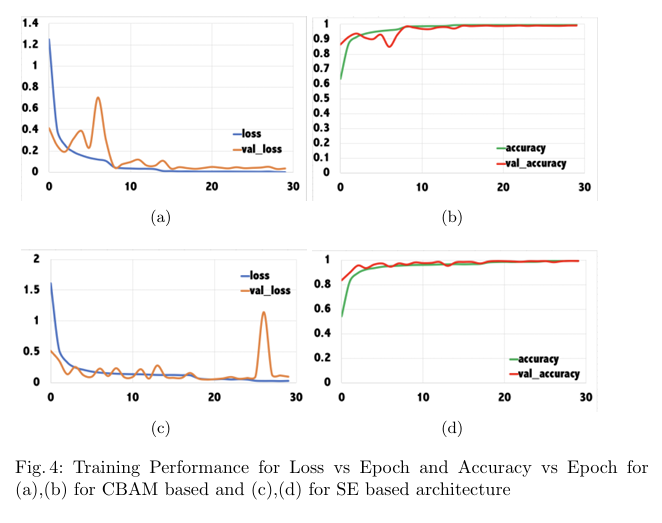
Plant diseases pose a significant threat to global food secu rity, demanding faster and more accessible methods for identification. While deep learning has shown promise in automating plant disease diagnosis from digital images, large model sizes remain a hurdle. This paper proposes a novel lightweight deep learning framework based on CovXNet, a convolutional neural network architecture utilizing efficient depthwise convolutions, combined with Squeeze-and-Excitation (SE) and Convolutional Block Attention Module (CBAM) attention mechanisms for enhanced performance. Our proposed models have been evaluated on the publicly available Plant Village dataset, achieving state-of-the art performance with 99.37% test accuracy using CovXNet with SE and 99.30% using CovXNet with CBAM across 38 classes. These re sults demonstrate the effectiveness of our approach in facilitating accu rate and efficient plant disease diagnosis, particularly for deployment on resource-constrained devices.
BibTeX
Click to copy@inproceedings{chowdhury2024attention,
title={Attention-Enhanced Multi-dilation CNN for Plant Disease Classification},
author={Chowdhury, Disha and Bashree, Nomaiya and Ovi, Tareque Bashar and Nyeem, Hussain and Wahed, Md Abdul and Tanzim, Rawnak},
booktitle={International Conference on Human-Centric Smart Computing},
pages={111--121},
year={2024},
organization={Springer}
}
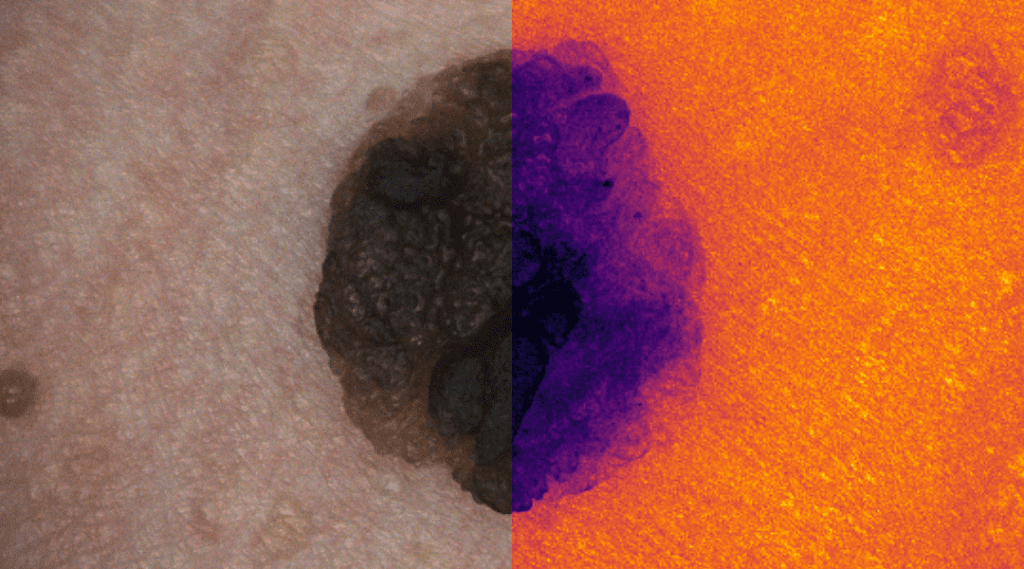
Skin Cancer Detection
Deep learning for skin cancer relies on segmentation and classification.
This research area focuses on developing advanced deep learning frameworks for the automated detection, segmentation, and classification of skin lesions from dermoscopic and clinical images. By leveraging convolutional neural networks (CNNs) and transformer-based architectures, the goal is to differentiate malignant melanomas from benign skin conditions with high diagnostic accuracy. The research integrates explainable AI techniques to enhance clinical trust and decision support, ultimately contributing to early detection and improved patient outcomes in dermatological diagnostics.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos
No photos added.
Ongoing Research Projects
Objective
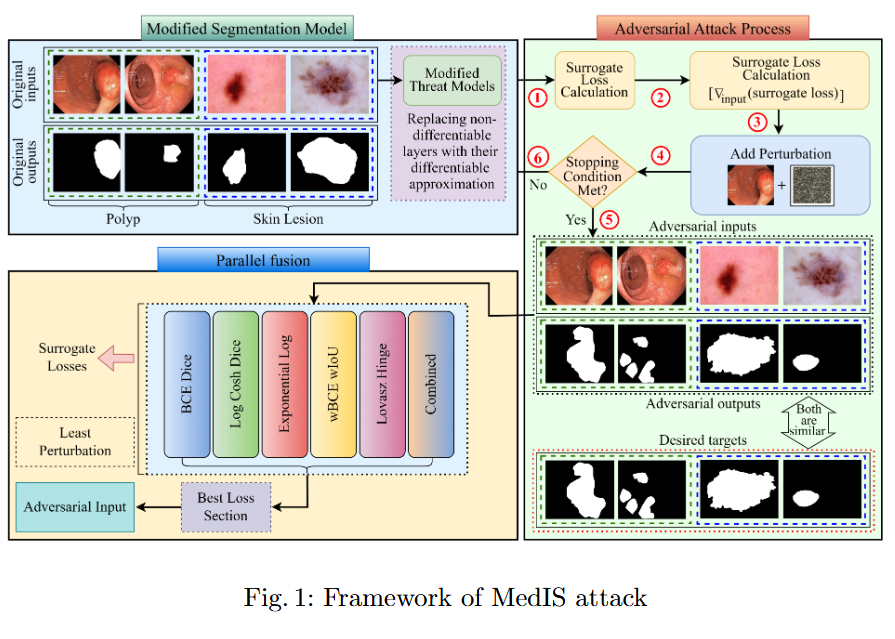
This study evaluates the robustness of leading segmentation models against advanced adversarial attacks in the context of polyp and skin lesion identification.
Key Findings
- We benchmark the robustness of leading segmentation models under strong adversarial attacks, systematically identifying their failure modes in the context of polyp and skin lesion tasks.
- The first systematic study of bi-level nested architectures under adversarial threat, clarifying how multiscale fusion and cross-level aggregation influence resilience.
- Evidence that global feature representations mitigate adversarial perturbations, with distilled design guidelines for more robust medical segmentation systems.
Photos
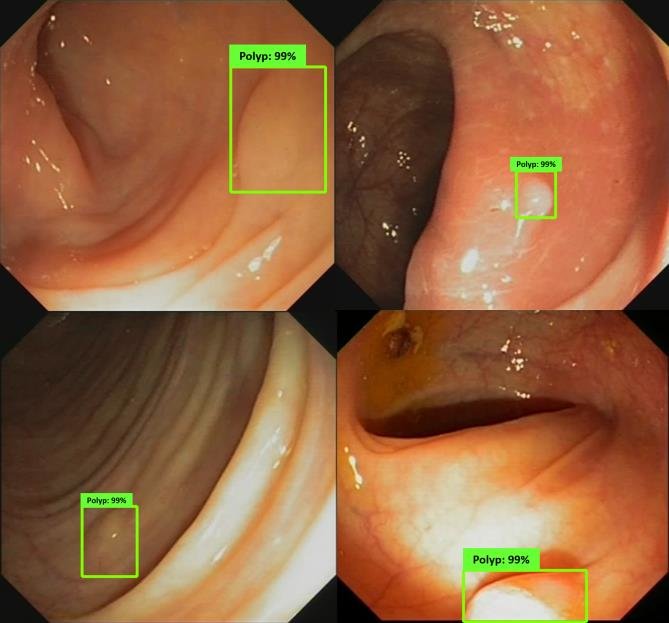
Colonoscopic Polyp Detetction
An AI "second eye" for real-time polyp detection during colonoscopy.
Research in colonoscopic polyp detection aims to design real-time AI-assisted systems for the early identification of colorectal polyps during colonoscopy. Using deep convolutional and attention-based segmentation models, the objective is to improve the sensitivity and precision of polyp localization under varying illumination, motion, and texture conditions. The developed models function as a “second observer” for endoscopists, assisting in diagnostic decisions and reducing the rate of missed polyps, thereby enhancing colorectal cancer prevention.
Selected Research Projects
No selected projects in this area.
Ongoing Research Projects
Objective
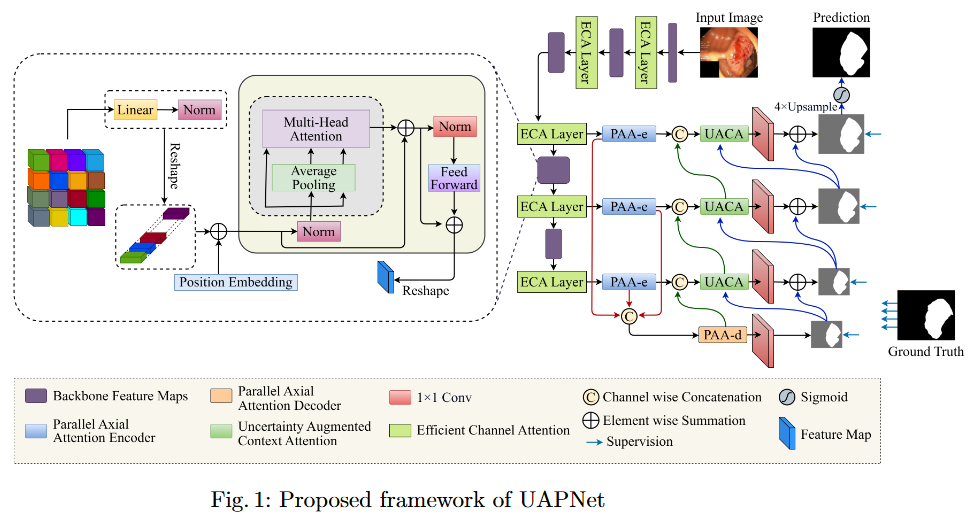
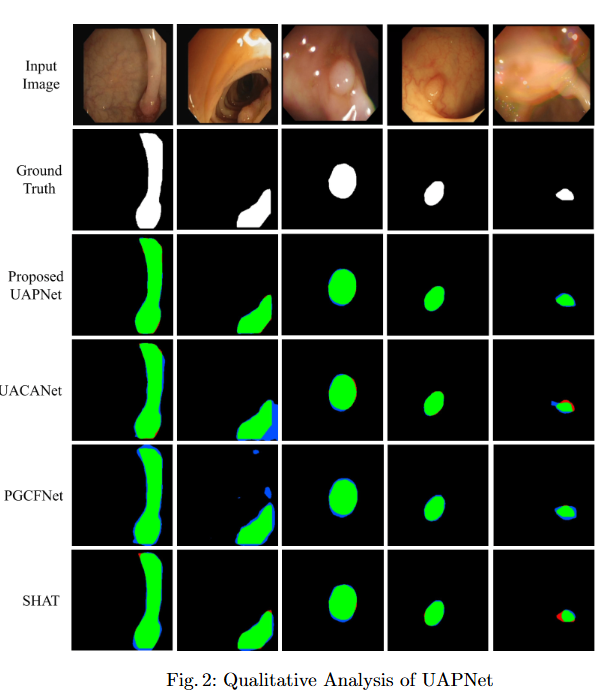
To mitigate feature redundancy in deep networks, we integrate the Efficient Channel Attention (ECA) module with the PVT encoder and augment it with an Uncertainty Augmented Context Attention (UACA) mechanism.
Key Findings
- PVT with Uncertainty-Augmented Context Attention. We integrate the PVT encoder with an uncertainty-augmented context attention mechanism, enabling the model to focus explicitly on ambiguous boundary regions. This mechanism improves the accuracy of boundary delineation by using uncertainty as a guiding signal.
- Integrating ECA with PVT Based Encoder. We incorporate ECA after each PVT stage, which refines feature representations while maintaining the original dimensionality. This enhancement improves the model’s discriminative power without introducing additional computational overhead.
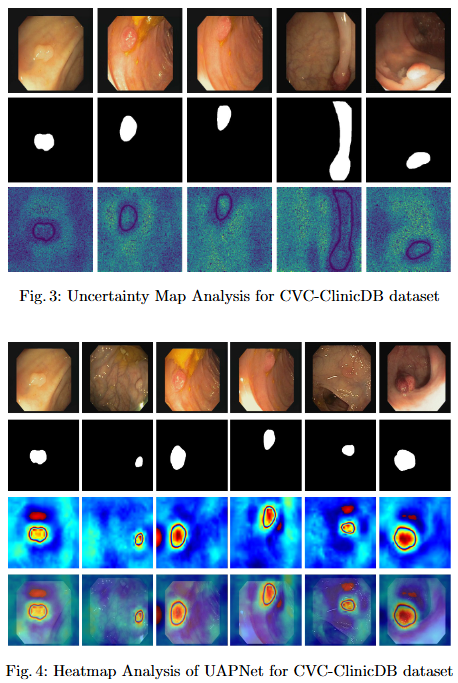
- Explainability for Clinical Reliability. To ensure transparency and clinical applicability, we apply explainable AI (XAI) techniques, including uncertainty maps and heatmaps for foreground and edge regions. These methods improve interpretability and provide clinicians with more reliable decision making tools by highlighting areas of uncertainty and focus during polyp segmentation.
Photos
Objective
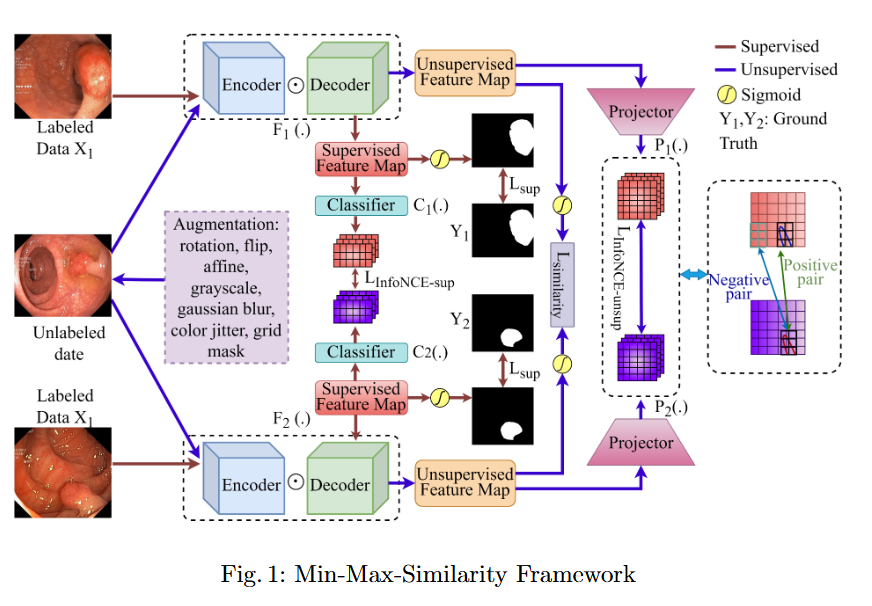
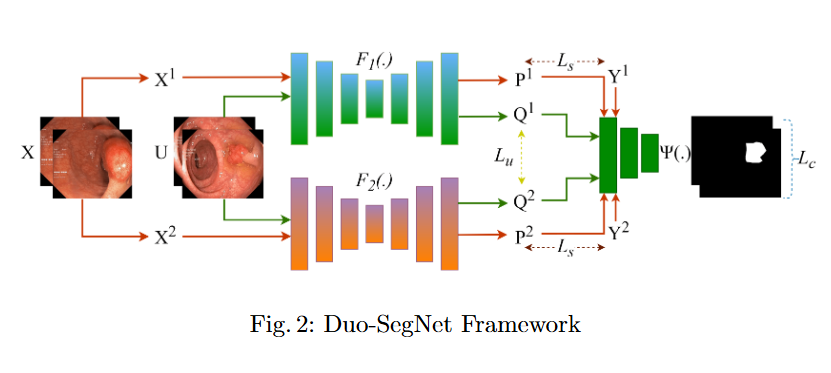
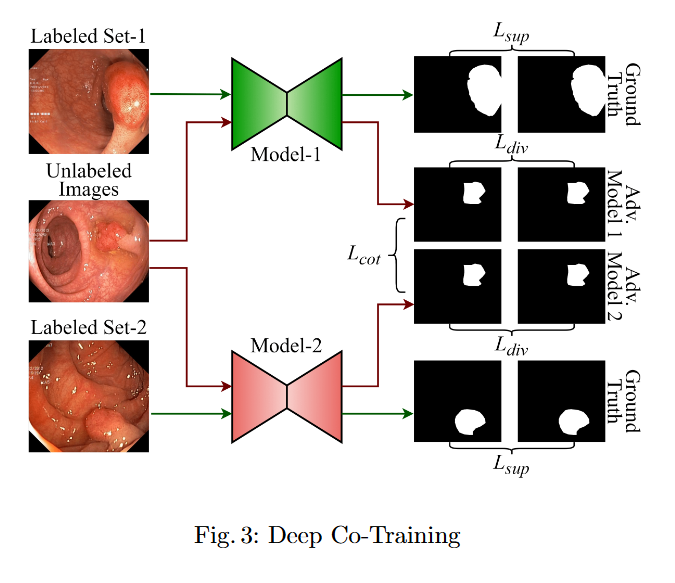
In this study, we systematically investigate how established supervised segmentation models perform within three representative semi-supervised frameworks: Min-Max Similarity, Duo-SegNet, and Deep Co-Training.
Key Findings
- We represent a comprehensive evaluation of multiple supervised segmentation architectures in semi-supervised learning scenarios.
- We identify model characteristics that enable effective utilization of unlabeled data within semi-supervised frameworks.
- We provide empirical insights that inform the design of more better and dependable semi-supervised polyp segmentation methods, thereby reducing annotation requirements while maintaining high segmentation accuracy.
Photos
Objective
This study evaluates the robustness of leading segmentation models against advanced adversarial attacks in the context of polyp and skin lesion identification.
Key Findings
- We benchmark the robustness of leading segmentation models under strong adversarial attacks, systematically identifying their failure modes in the context of polyp and skin lesion tasks.
- The first systematic study of bi-level nested architectures under adversarial threat, clarifying how multiscale fusion and cross-level aggregation influence resilience.
- Evidence that global feature representations mitigate adversarial perturbations, with distilled design guidelines for more robust medical segmentation systems.
Photos

Medical Image Analysis
Imaging, segmentation, and diagnostics for medical data.
This area encompasses a broad spectrum of imaging modalities—MRI, CT, X-ray, and endoscopic imaging—to develop computational tools for segmentation, registration, and diagnostic interpretation. Research focuses on integrating multimodal image features with deep learning to improve disease localization and treatment planning. The emphasis is on constructing efficient, explainable, and data-driven models that support clinical workflows, thereby bridging the gap between medical imaging research and practical healthcare applications.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos




Objective
No objective provided.
Key Findings
No findings listed yet.
Photos
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos
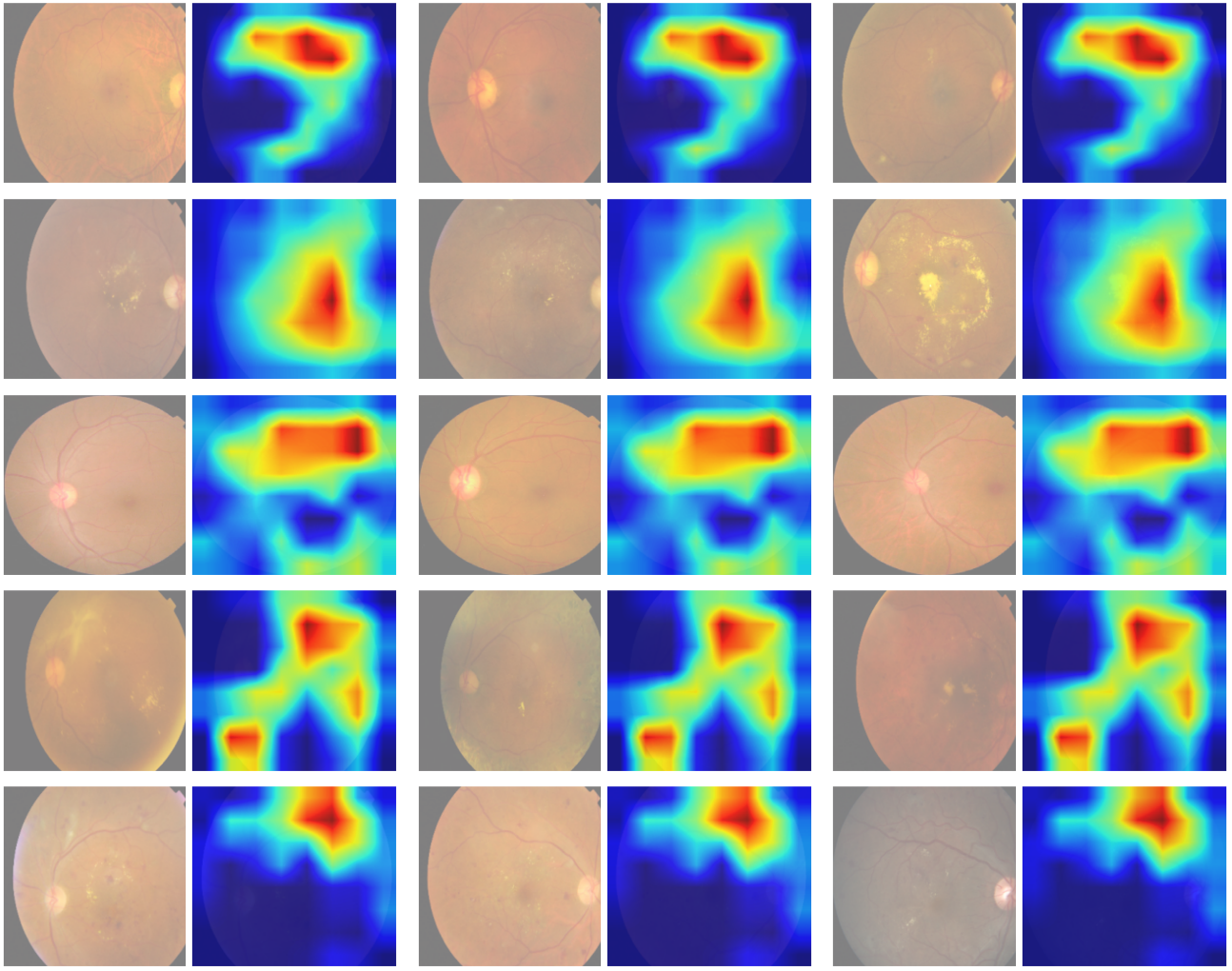
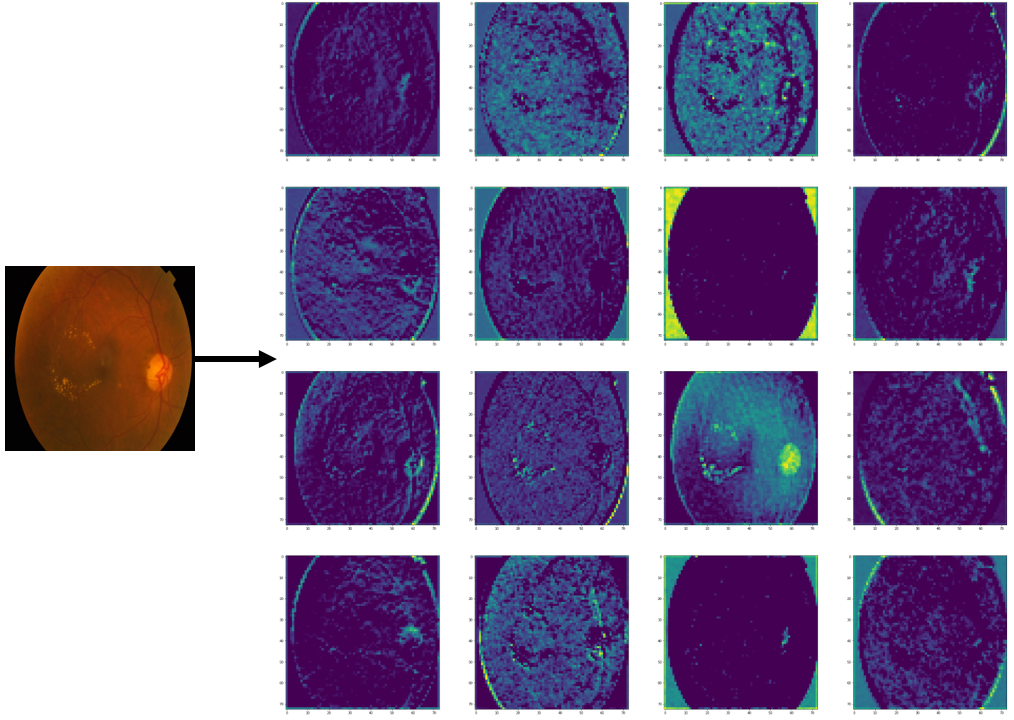
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos
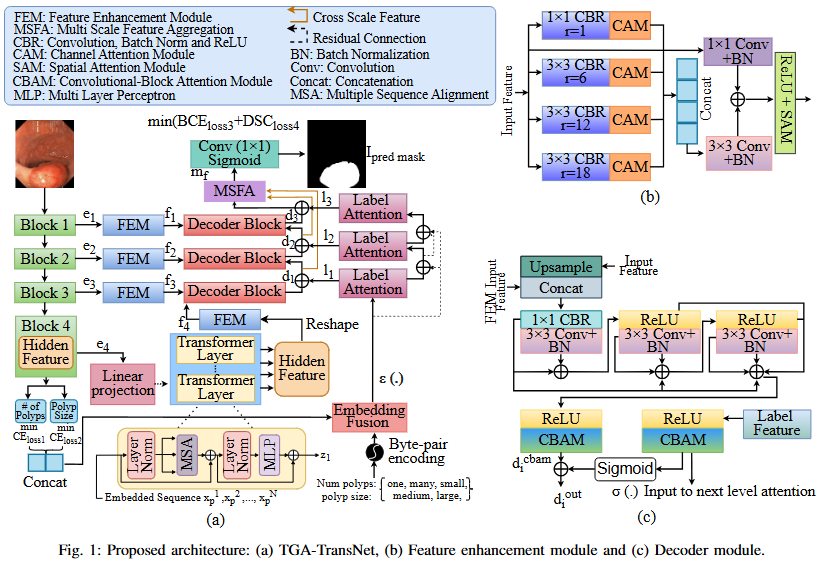
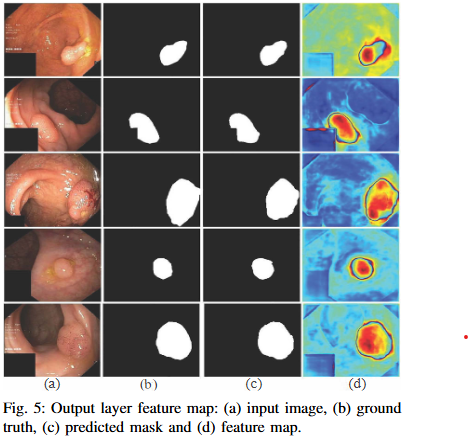
Ongoing Research Projects
No ongoing projects in this area.

Computer Vision
Algorithms and systems for visual understanding and perception.
Computer vision research at the lab explores the development of algorithms and systems capable of perceiving and understanding visual data in real-world environments. Topics include object detection, scene understanding, tracking, and visual reasoning. By integrating attention mechanisms, transformers, and graph neural networks, this work advances autonomous perception systems across domains such as healthcare, robotics, and intelligent surveillance, pushing the boundaries of visual cognition and artificial intelligence.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos


Objective
No objective provided.
Key Findings
No findings listed yet.
Photos

Objective
No objective provided.
Key Findings
No findings listed yet.
Photos




Ongoing Research Projects
No ongoing projects in this area.

Remote Sensing
Earth observation and geospatial analysis from aerial and satellite imagery.
This research investigates the use of aerial and satellite imagery for earth observation and geospatial analysis. Deep neural networks are employed for tasks such as land cover classification, road and building extraction, and environmental monitoring. The goal is to develop accurate and scalable models capable of processing high-resolution satellite data, thereby supporting urban planning, disaster management, and sustainable resource monitoring with precision-driven remote sensing analytics.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos



Objective
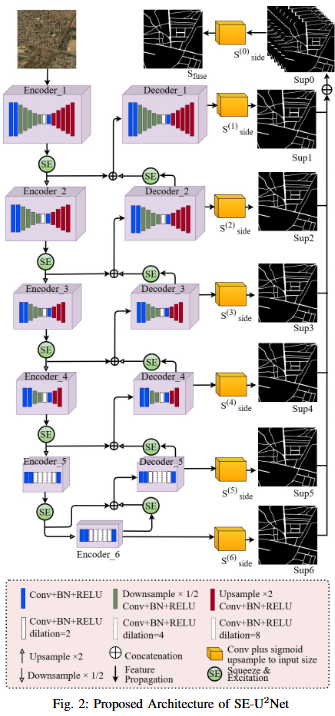
To develop SE-U2Net, a robust and efficient deep learning framework for road extraction from very high-resolution (VHR) satellite imagery by integrating the U²-Net architecture with Squeeze-and-Excitation (SE) modules, enabling selective channel refinement and multi-scale contextual feature learning without reliance on pre-trained backbones.
Key Findings
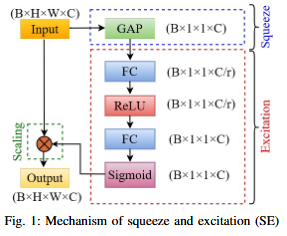
- Architectural Innovation: Integration of Squeeze-and-Excitation (SE) blocks for dynamic channel feature recalibration.
- Performance Improvement: Enhanced discrimination of minor roads through learned channel interdependencies.
- Computational Efficiency: Parameter-efficient attention mechanism achieved without complex spatial operations.
Photos
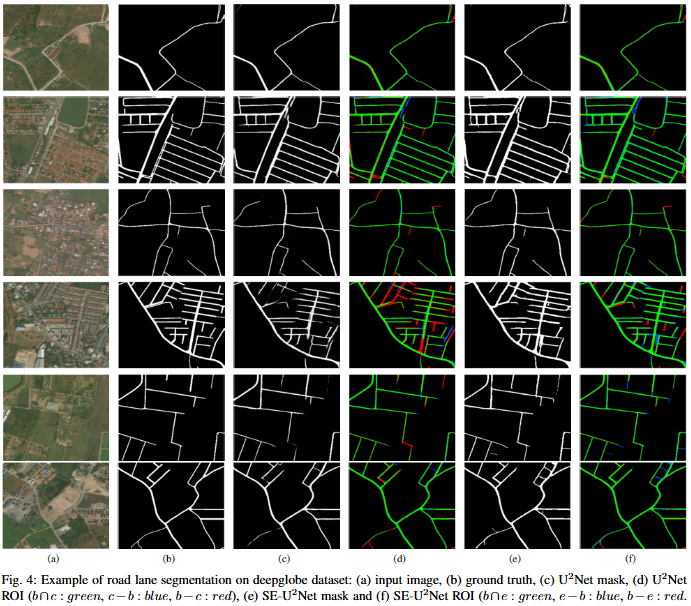

Objective
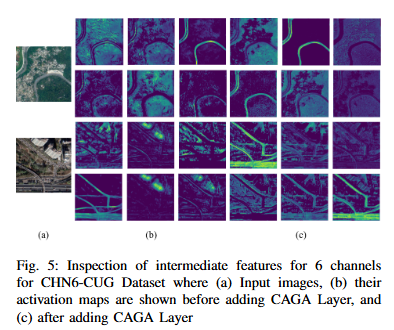
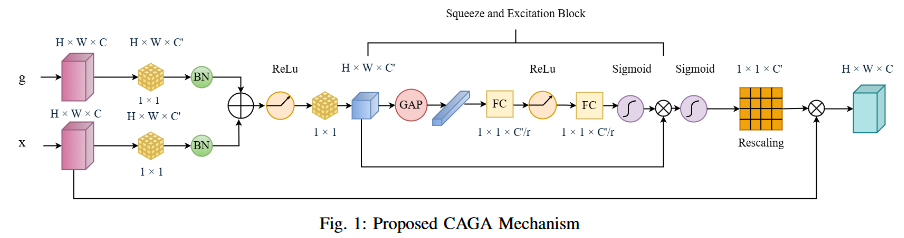
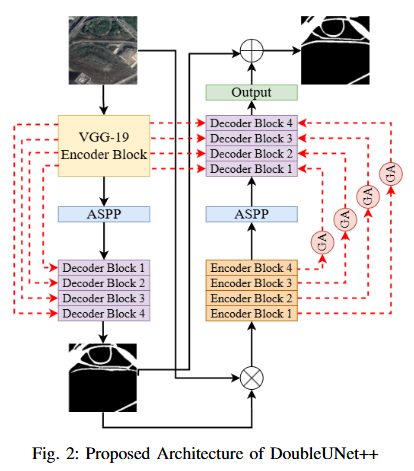
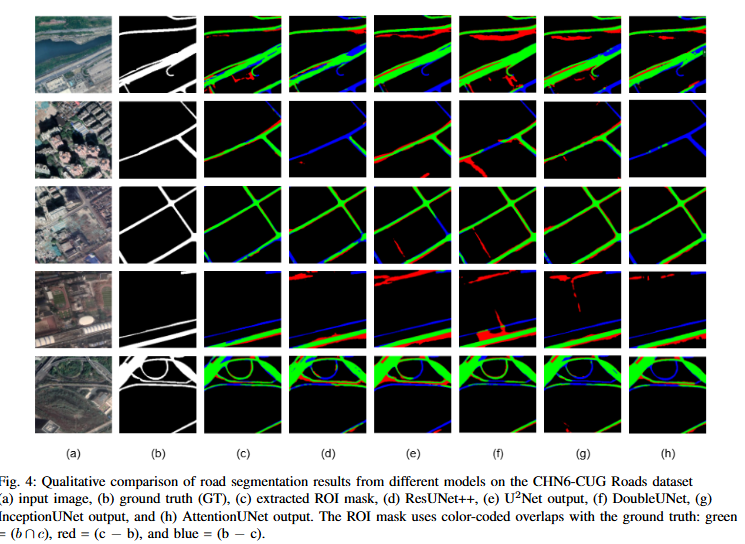
To develop DoubleUNet++, an enhanced road extraction model that improves the contextual understanding and spatial precision of remote sensing image segmentation by integrating a Channel-Aware Gated Attention (CAGA) mechanism and Squeeze-and-Excitation (SE) layers into the DoubleUNet architecture.
Key Findings
- We propose Channel-Aware Gated Attention (CAGA) mechanism which has been integrated into skip connections of the secondary U-Net, enabling dynamic spatial feature weighting to emphasize critical road regions while suppressing irrelevant details
- A squeeze-and-excitation (SE) layer preceding the final activation in the attention pathway, adaptively recalibrating channel-wise feature responses to enhance discriminative power
Photos

Ongoing Research Projects
No ongoing projects in this area.

Computational Biology and Bioinformatics
Data-driven modeling and analysis of biological systems.
The lab’s research in computational biology and bioinformatics applies machine learning and statistical modeling to the analysis of complex biological data. Efforts include protein structure prediction, gene expression analysis, and biological network modeling. Through the integration of omics data and imaging modalities, this area aims to uncover biological patterns and enhance understanding of disease mechanisms, ultimately contributing to precision medicine and drug discovery initiatives.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos




Ongoing Research Projects
Objective
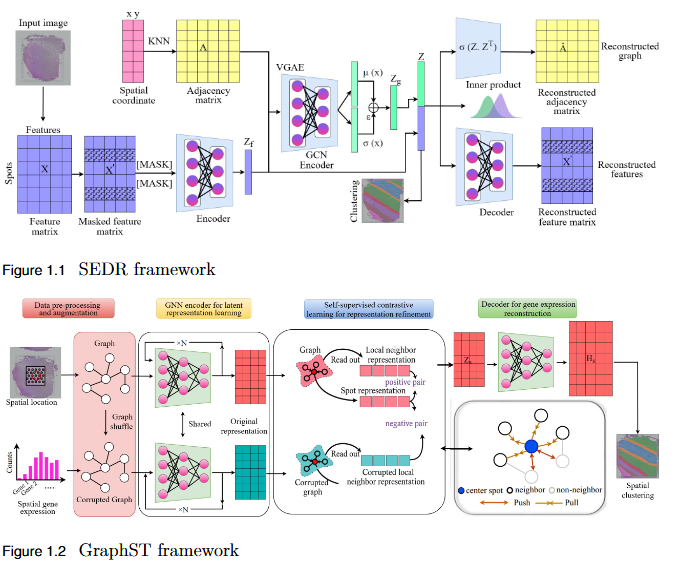
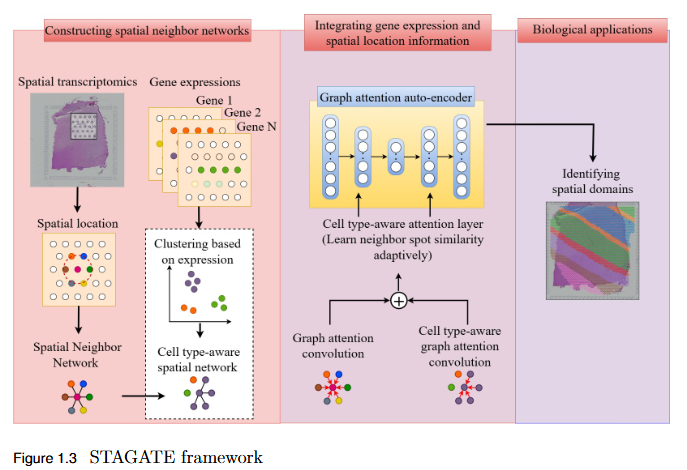
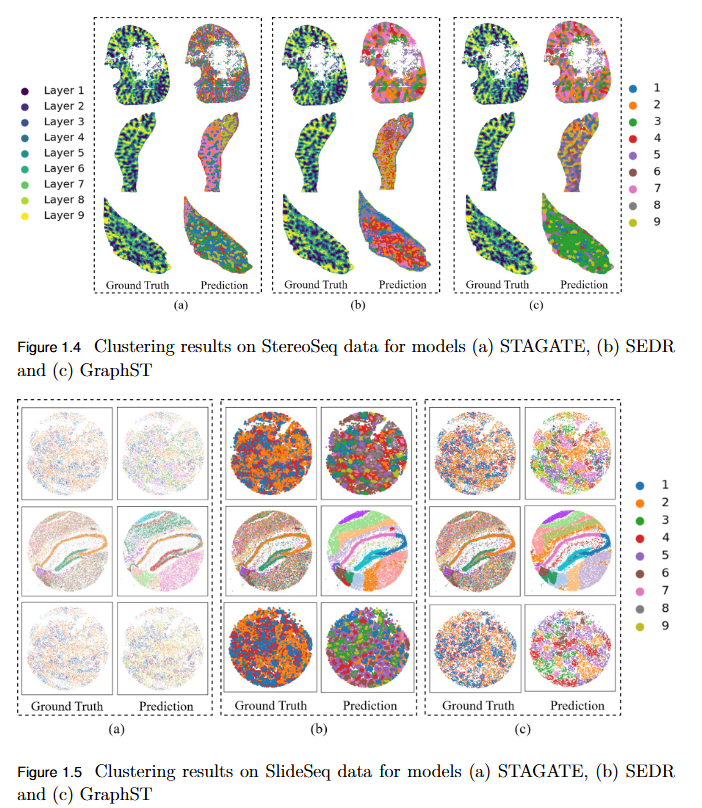
To systematically evaluate the performance and scalability of leading GNN-based spatial transcriptomics (ST) models—STAGATE, GraphST, and SEDR—on high-density next-generation datasets such as Stereo-seq and Slide-seq, and to identify their limitations in modeling complex spatial structures.
Key Findings
- Performance Evaluation: Comprehensive assessment using ARI, AMI, NMI, and HOMO metrics revealed modest and variable clustering performance across high-density datasets.
- - SEDR achieved the highest scores on Slide-seq datasets, but overall accuracy remained low.
- - Stereo-seq results were inconsistent, with no single model consistently outperforming others.
- Scalability Limitation: Existing GNN-based ST models struggle to generalize to high-resolution data, indicating poor scalability to large, complex tissue structures.
- Benchmark Establishment: This study provides a performance baseline for evaluating future models on next-generation ST platforms.
- Research Implication: Highlights the urgent need for developing new computational frameworks that are more robust, adaptive, and scalable for analyzing dense spatial transcriptomics data.
Photos
Objective
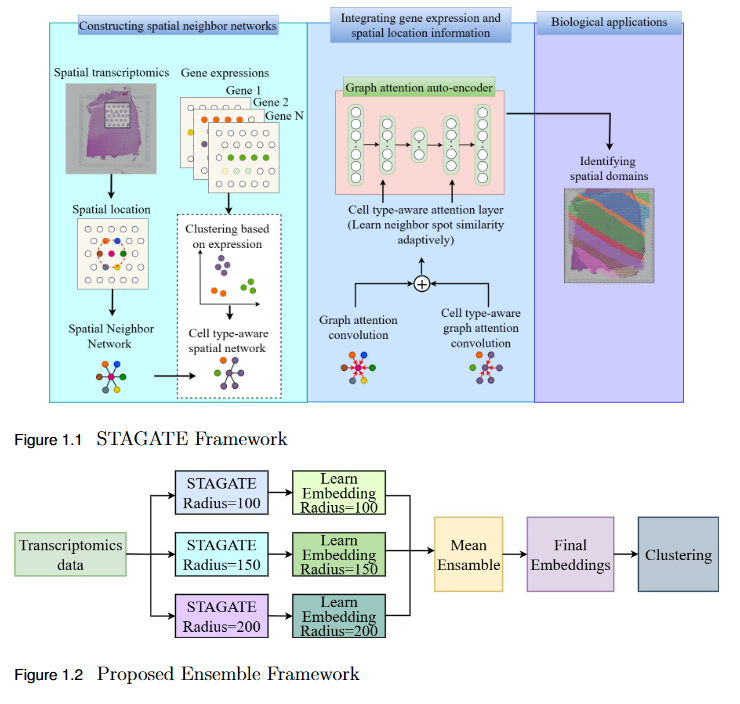
To overcome the limitation of fixed spatial neighborhood scales in existing spatial transcriptomics (ST) models by developing En-STAGATE, an ensemble framework that integrates multiple spatial graphs across varying neighborhood radii to capture both local and global tissue structures for improved domain identification.
Key Findings
- Multi-Scale Ensemble Design: En-STAGATE constructs and fuses representations from multiple spatial graphs at different neighborhood radii, effectively modeling both fine-grained cellular interactions and large-scale tissue organization.
- Enhanced Biological Interpretability: The multi-scale integration provides a more comprehensive and biologically meaningful latent representation of tissues.
- Empirical Validation: Achieved competitive or superior performance across 14 benchmark datasets, outperforming STAGATE and other leading methods in several key cases.
- Improved Domain Delineation: Notably higher Adjusted Rand Index (ARI) scores on datasets such as 151674, 151675, 151676, and Mouse, demonstrating robustness in complex tissue structures.
- Key Insight: Highlights that a single neighborhood scale is insufficient for spatial domain identification, establishing multi-scale analysis as a critical paradigm in spatial transcriptomics.
Photos
Objective
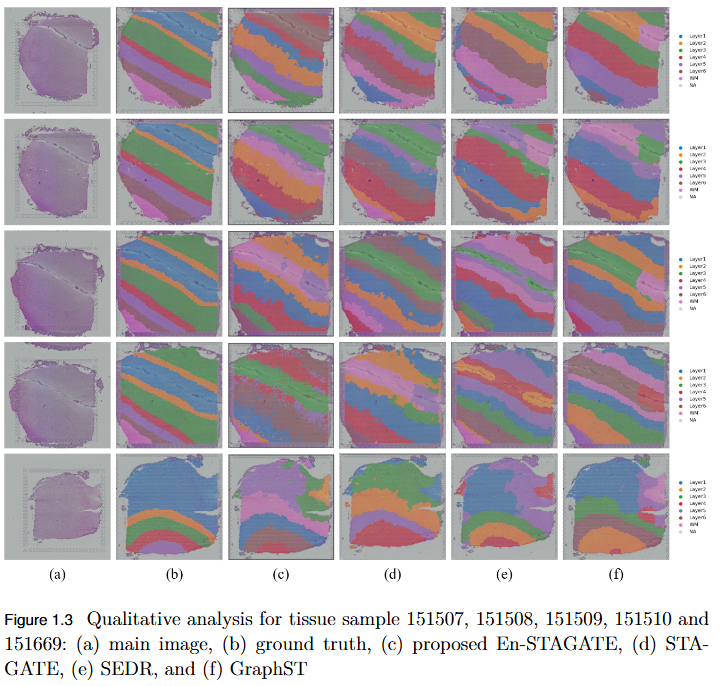
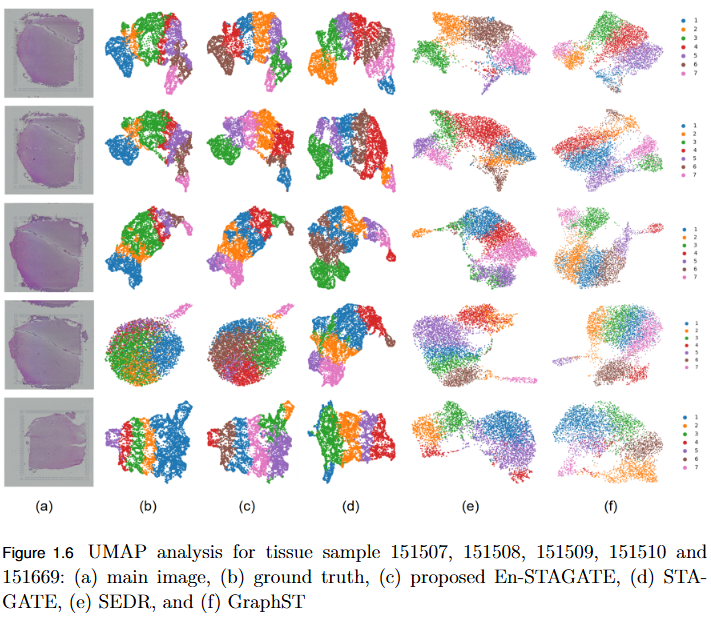
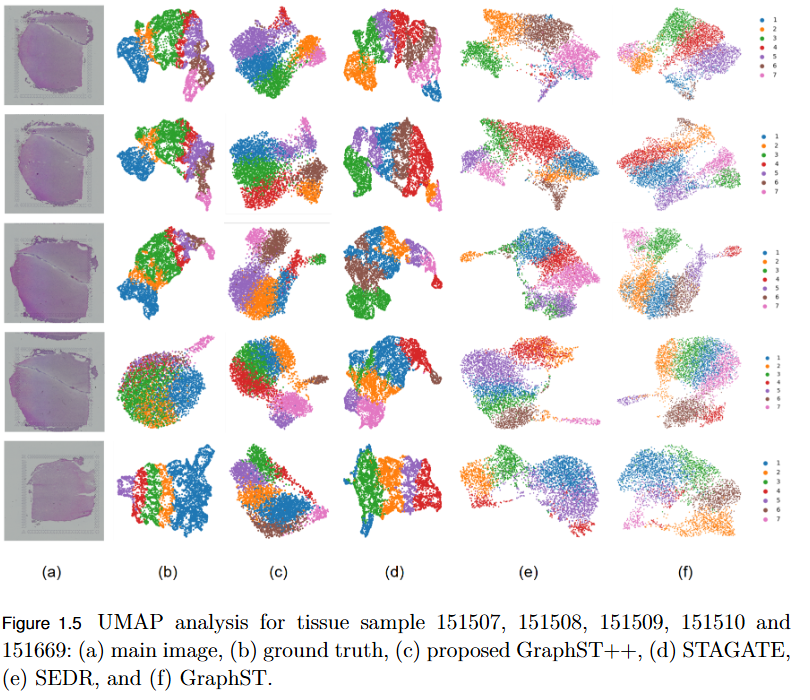
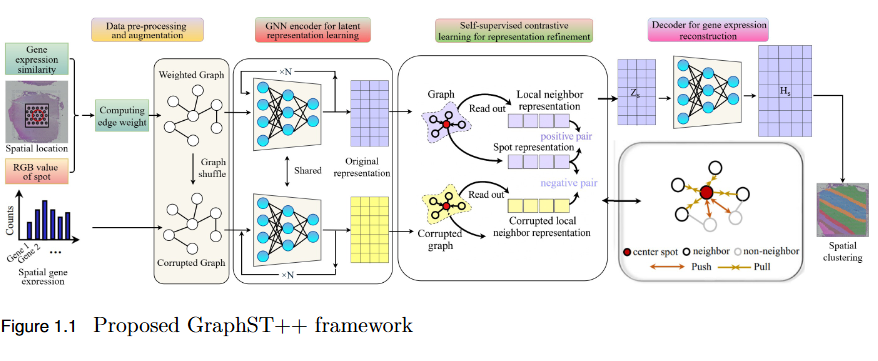
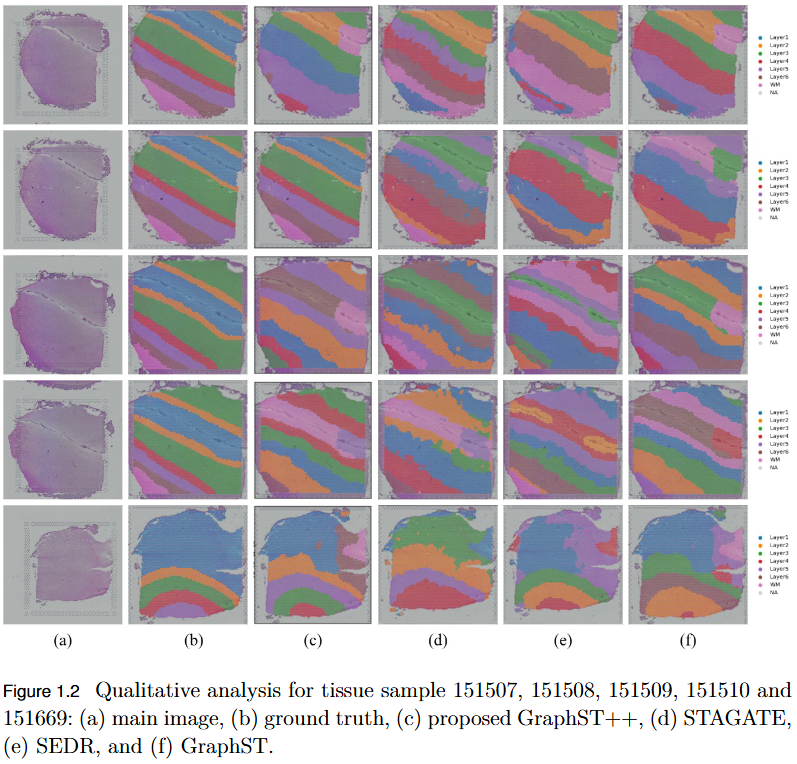
To address the over-smoothing and spatial bias in current spatial transcriptomics (ST) models by developing GraphST++, a multi-modal framework that integrates spatial, gene expression, and histological features for constructing biologically faithful tissue graphs.
Key Findings
- Multi-Modal Graph Construction: GraphST++ defines inter-spot relationships using a composite score combining spatial distance, gene expression similarity, and histological morphology, reducing over-reliance on proximity-based connections.
- Improved Tissue Representation: The proposed approach captures complex biological interactions and produces a more accurate reflection of the tissue microenvironment.
- Superior Performance: Demonstrated higher Adjusted Rand Index (ARI) compared to leading models, including GraphST, across multiple benchmark datasets such as human breast cancer (BRCA) and human brain tissues.
- Robustness Across Datasets: While not universally dominant, GraphST++ consistently benefits from morphological integration, yielding more stable and biologically interpretable clustering results.
- Broader Implications: Highlights the importance of multi-modal fusion in advancing spatial transcriptomics analysis and enabling more precise tissue domain identification.
Photos

Objective
To overcome the limitation of existing graph-based spatial transcriptomics (ST) models in capturing long-range spatial dependencies by developing STAGATE++, a hybrid GAT–Transformer architecture that jointly learns local and global tissue representations.
Key Findings
- Hybrid Architecture Advantage: STAGATE++ integrates a Graph Attention Network (GAT) for precise local feature extraction with a Transformer encoder for modeling long-range spatial dependencies.
- - Enhanced Spatial Awareness: The model constructs a holistic and spatially coherent tissue representation, improving understanding of tissue organization.
- Empirical Superiority: Demonstrated state-of-the-art performance across 14 benchmark datasets, consistently outperforming existing methods in spatial domain identification.
- Validation Metric: Significant improvements in Adjusted Rand Index (ARI) confirm the importance of global context modeling in tissue delineation.
- Future Extensions: Plans include incorporating histopathological image features, applying self-supervised training for better generalization, and integrating GNN interpretability techniques for biological insight and clinical transparency.
Photos
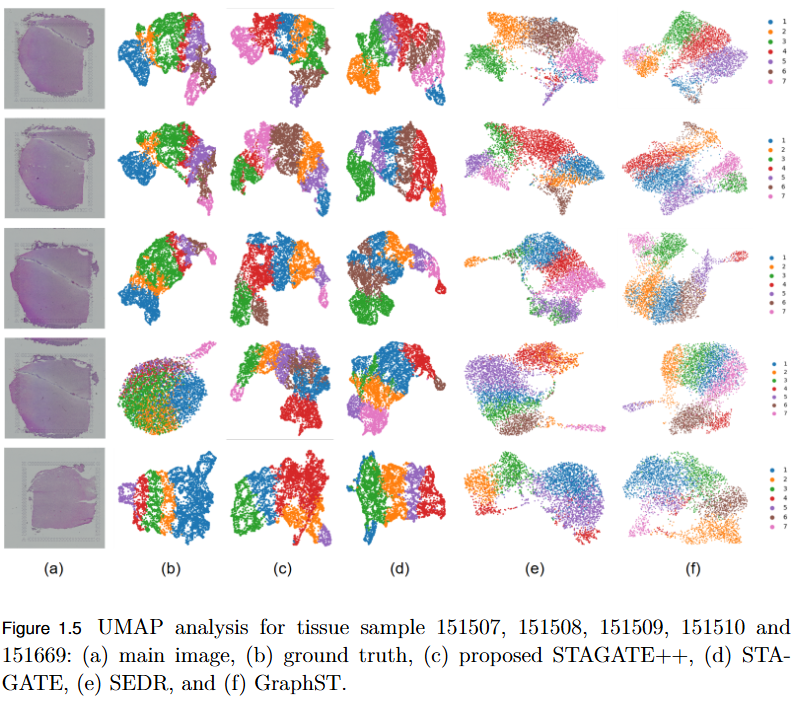
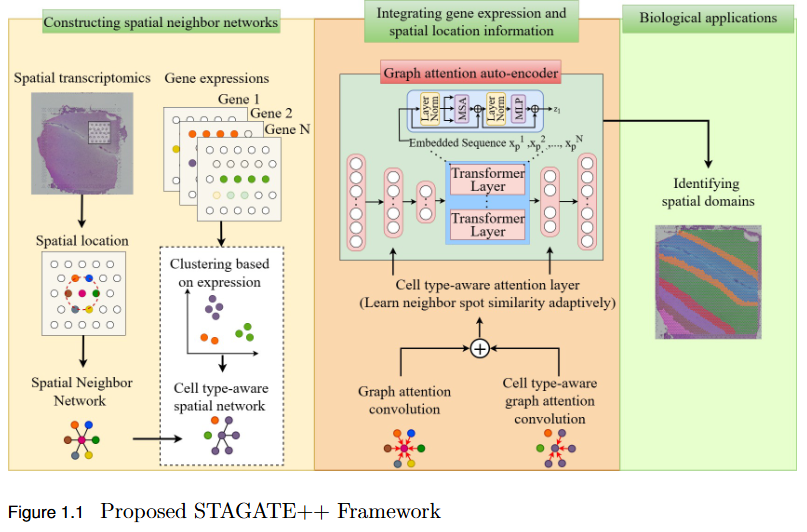
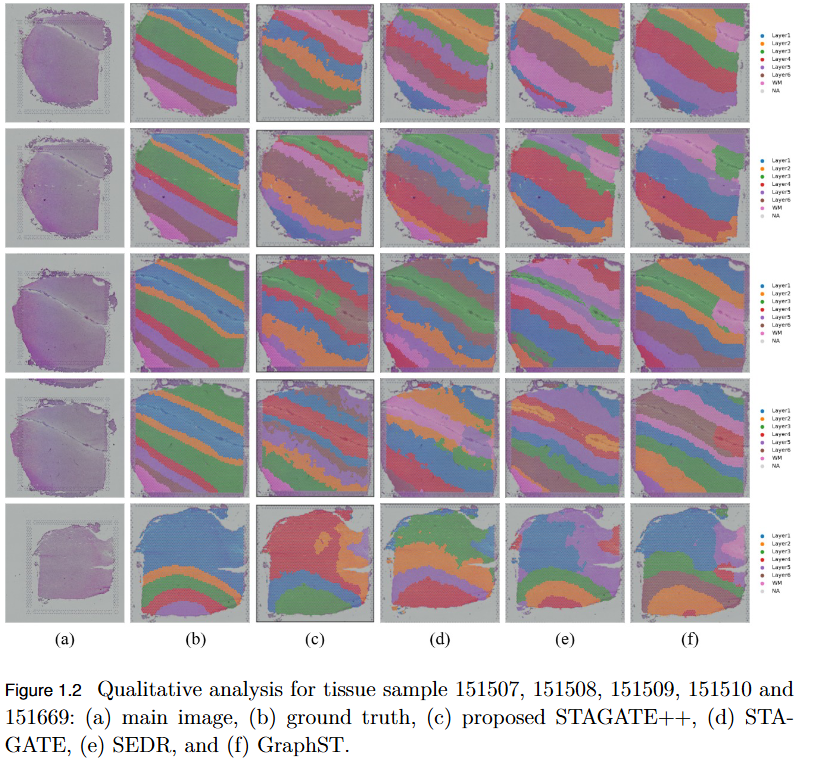


Image Processing
Signal and image enhancement, restoration, and transformation techniques.
This foundational area focuses on the theoretical and practical aspects of image enhancement, restoration, compression, and transformation. Research includes spatial and frequency-domain techniques for noise reduction, contrast enhancement, and feature extraction. By combining classical methods with modern deep learning frameworks, the objective is to achieve efficient, high-quality image processing pipelines that serve as the backbone for advanced applications in computer vision, medical imaging, and multimedia analysis.
Selected Research Projects
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos
Objective
No objective provided.
Key Findings
No findings listed yet.
Photos




Ongoing Research Projects
No ongoing projects in this area.
